 
The last step, individually analyzing each aptamer’s binding, kills any flickering hope of high-throughput aptamer discovery.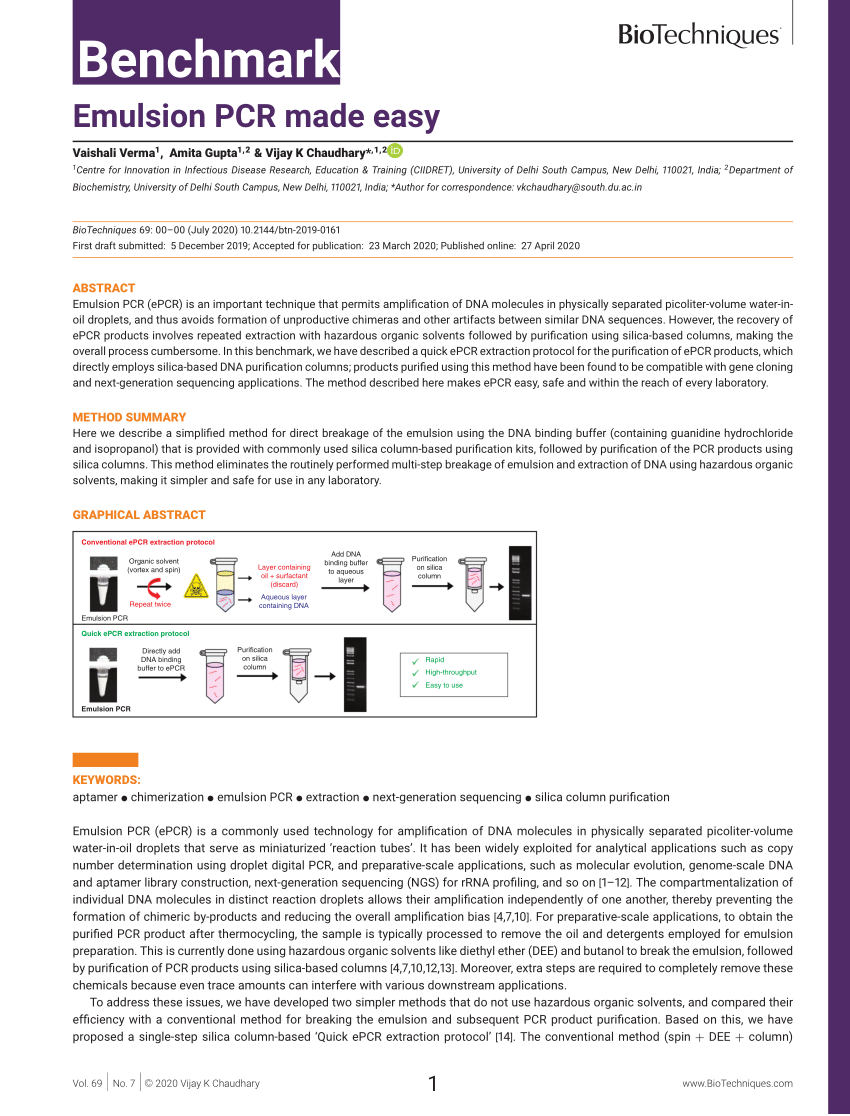 
These candidate aptamers are sequenced-an expensive step-and their sequences compared to identify aptamers that might bind well. It involves cloning candidate aptamer sequences into plasmids, transfecting these into bacteria, growing up the bacteria, and picking clones for aptamer analysis. After amplifying a library of possible sequences, actually generating the aptamers is quite time-consuming. Problem: Designing aptamers is generally a laborious, inefficient process. Aptamers that bind Shp2 and block its activity may have potential for treating Shp2-dependent cancers. Project: Efficiently select aptamers-segments of DNA with specific binding properties-against targets such as the cancer biomarker called SH2 domain–containing protein tyrosine phosphatase 2 (Shp2). Laboratory: Chaoyong James Yang, Lu Jiaxi Professor, Department of Chemical Biology, Xiamen University, Fujian, China Here are three case studies of researchers stretching ePCR’s capabilities. 
Tiemann-Boege and Novak are two of many scientists tweaking ePCR to maximize its benefits while minimizing its weaknesses. With standard PCR, you can amplify 1 to 2 kilobases,” Tiemann-Boege explains, but “the maximum in ePCR is 150 base pairs.” Commercial ePCR can also be a problem for anyone needing reaction volumes larger than a picoliter, prompting scientists to continue searching for creative solutions, says Richard Novak, of the University of California, Berkeley. In ePCR, “the product length is very short. Unfortunately, says Irene Tiemann-Boege, who studies genome recombination at Johannes Kepler University in Linz, Austria, emulsion PCR (ePCR) has a serious drawback for those hoping to cast a careful eye over the entire genome. (See “Prime Time for Digital PCR,” The Scientist, December 2011.) A plus is that the large number of individual reactions minimizes cross-contamination between products, but the greatest advantage is the ability to turn a tedious experiment into a high-throughput extravaganza. Our simple polydisperse droplet preparation and statistical framework can be extended to a variety of settings for the quantification of nucleic acids in complex samples.The amplification reaction can be carried out in many of the digital PCR machines on the market. We also report a multiplexed ddPCR assay and demonstrate proof-of-concept methods for rapid droplet preparation in multiple samples simultaneously. In this work, we show that these ddPCR assays can reduce overall assay time while still providing quantitative results. Additionally, this approach is compatible with a range of input sample volumes, extending the assay dynamic range beyond that of commercial ddPCR systems. The polydisperse droplet system allows for accurate quantification of droplet digital PCR (ddPCR) and reverse transcriptase droplet digital PCR (RT-ddPCR) that is comparable to commercially available systems such as BioRad's ddPCR. To address these limitations and make this technology more applicable for a variety of settings, we have developed a statistical framework to apply to droplet PCR performed in polydisperse droplets prepared without any specialized equipment. Though impactful, these improvements have generally been restricted to centralized laboratories with trained personnel and expensive equipment. These individual reaction vessels lead to digitization of PCR enabling improved time to detection and direct quantification of nucleic acids without a standard curve, therefore simplifying assay analysis. Lab-on-a-chip applications have developed methods to partition single biomolecules, such as DNA and RNA, into picoliter-sized droplets. Nucleic acid amplification technology, such as polymerase chain reaction (PCR), has enabled highly sensitive and specific disease detection and quantification, leading to more accurate diagnosis and treatment regimens. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
